Your cart is currently empty!
Proteomics
Harness the power of proteomics.
Explore the power of data independent acquisition (DIA), also known as SWATH® Acquisition. A label-free quantification approach is designed to achieve optimal quantitative data analysis of several thousand proteins in various sample types.

BEst-in-Class Quantification
Label-free proteomic quantification
Quantify every single peptide
Our team of scientists has optimized the label-free quantitative proteomics solution to provide the most reliable result for your workflow.
Using SWATH acquisition which is also called data independent acquisition (DIA) method, every single ionizable peptide in the sample can be detected and quantified. The method is ideal for comparison studies between cohorts of two or multiple different treatments.
Divide to see better
The SWATH technology uses a clever way to record everything while maintaining the ability to easily search the results after the analysis. The key to this acquisition method is a fractionation of the studied mass range in smaller windows. Every peptide that falls into one window is fragmented and the machine records its MS/MS signature. Then, by using an ion library, we are able to deconvolute the signal and extract quantitative information for every peptide, at the MS/MS level.
Profiling Proteomics
Multiplex your samples to profile thousands of proteins in a single analysis
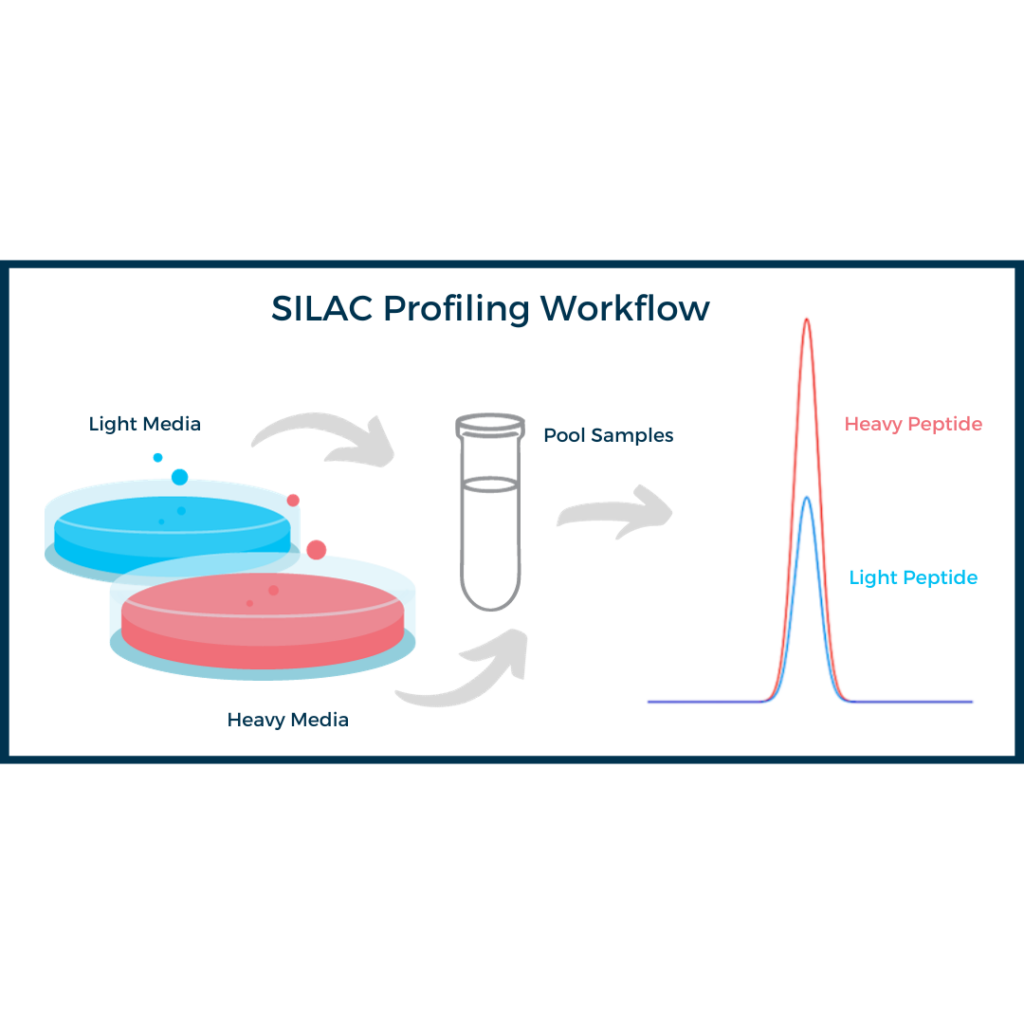
Sample multiplexing, i.e. pooling several experimental conditions in one MS run, is a method that helps with proteome profiling.
Labeling techniques provide the highest level of comparison between the treatment groups. Differential chemical labeling allows sample multiplexing which provides powerful tools for proteome profiling without extensive fractionation of samples before analysis.
Some examples of labeling techniques are:
- TMT
- SILAC (stable isotope labeling with amino acids in cell culture)
- iTRAQ (isobaric amine-specific labelling, up to 8plex)
- mTRAQ (non-isobaric amine specific labelling, up to 3plex)
- Cysteine-targeted labelling
- Dimethyl labeling
Data Analytics
We help our customers make data-driven progress.
Our team transforms your raw data in to meaningful insights. Nothing gets lost in translation.
Our in-depth and interactive data reports include analyzed data, methods, and conclusions. You can also access your raw data. We don’t hold anything back.
It’s not just a report. We connect with you to ensure the results are clear, help you gather outputs for key stakeholders, and plan next steps in your path to market.
- Actionable insights
- Interactive reports
- Advanced stats
- Visualization tools
- AI/ML-ready data
- Collaborative approach
Related Capabilities
Lipidomics
In-depth profiling of many classes of biologically relevant lipids using our high resolution instruments.
Multiomics
See the big picture by combining proteomics, lipidomics and metabolomics data.
Metabolomics
Sample profiling using either untargeted metabolomics or selected panels of targeted metabolites is the best technique.
Data Analytics
We help customers unlock the value and potential of their data with clearer, deeper insights.
Let’s connect
Get in touch with our experts
